Arbeitsgruppen am Physik-Department
Biomedizinische Physik (Prof. Franz Pfeiffer)
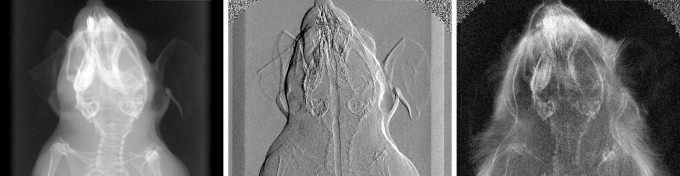
Our interdisciplinary research portfolio is focused on the translation of modern x-ray physics concepts to biomedical sciences and clinical applications. We are particularly interested in advancing conceptually new approaches for biomedical x-ray imaging and therapy. From a medical perspective, our work currently targets early cancer and osteoporosis diagnostics.
Biomolekulare Nanotechnologie (Prof. Hendrik Dietz)

We develop novel scientific devices and methods for applications in biomolecular physics, biological chemistry, and molecular medicine. To this end, we currently focus on using DNA as a programmable construction material for building nanometer-scale scientific devices with atomically precise features. We also customize proteins and create and study hybrid DNA-protein complexes. 3D transmission electron microscopy, atomic force microscopy, and single molecule methods such as optical trapping and fluorescence microscopy are among our routine analysis tools.
Molekulardynamik (Prof. Martin Zacharias)
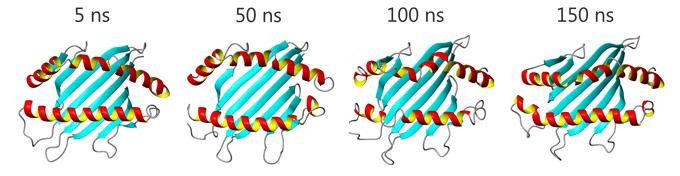
Die Funktion von Proteinen und Nukleinsäuren in lebenden Zellen wird durch ihre molekulare Dynamik bestimmt. Am Lehrstuhl „Molekulardynamik“ werden Computersimulationsmethoden eingesetzt, um die Struktur, Funktion und Dynamik von Biomolekülen zu untersuchen. Zu unseren Zielen zählt dabei ein genaueres Verständnis von Strukturbildungsprozessen und die Entschlüsselung des Mechanismus der Assoziation von Biomolekülen.
Molekulare Biophysik (Prof. Matthias Rief)
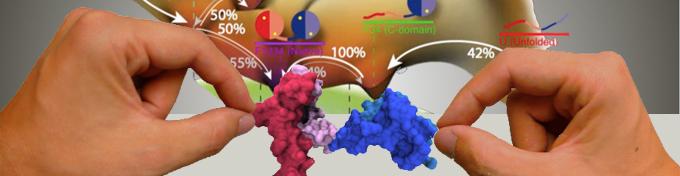
Proteins are fascinating examples of self-organized molecular machines. Without any help a polypeptide strand can fold into functional threedimensional structures. We are interested in studying the function and folding process of proteins on the single molecule level. Examples are single molecule folding/unfolding studies or the motility of molecular motors in optical traps.
Physik der biomedizinischen Bildgebung (Prof. Julia Herzen)
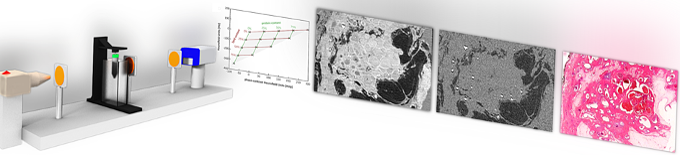
We develop novel X-ray imaging methods using highly brilliant synchrotron radiation and conventional laboratory X-ray sources. We mainly focus on quantitative multi-modal approaches combining spectral and phase-contrast imaging. Currently, we are aiming at applying these methods for improved breast cancer detection and for quantitative 3D virtual histology of human tissue.
Physik synthetischer Biosysteme (Prof. Friedrich Simmel)

Our goal is the realization of self-organizing molecular systems that are able to respond to their environment, compute, move, take action.
Theoretische Biophysik neuronaler Informationsverarbeitung
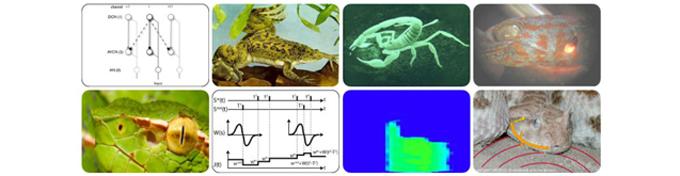

Wir erforschen Aufbau und Funktionsweise von informationsverarbeitenden Systemen in der Natur. Ziel ist es, aus grundlegenden Bausteinen der Wahrnehmung universelle Prinzipien abzuleiten. Ein solcher Baustein kann sowohl ein spezielles sensorisches System als Ganzes, als auch ein einzelnes Glied in einer sensorischen Verarbeitungskette repräsentieren. Dabei reduzieren wir die Prozesse der Informationsaufbereitung derart, dass mathematisches Verständnis und gleichzeitig experimentelle Überprüfbarkeit möglich werden.
Theorie biologischer Netzwerke (Prof. Karen Alim)
Wie entstehen Form und Struktur, wenn Leben entsteht? Wir wollen herausfinden, welche physikalischen Grundlagen für das Formen des Lebens verantwortlich sind. Um dies zu erreichen, kombinieren wir quantitative Beobachtung des Lebens mit theoretischen Modellen, um schließlich die Physik der Schlüsselprozesse in einfachen mathematischen Begriffen zu erfassen. Das Leben verbirgt noch immer viele grundlegende Prozesse, die, sobald sie aufgedeckt werden, unser Design, unsere Technik oder unsere medizinische Behandlung revolutionieren können.
Theorie komplexer Biosysteme (Prof. Ulrich Gerland)

In physics, interactions between particles follow laws. In biology, interactions between biomolecules serve a function. We study the physics of biological functions. In particular, we are interested in cases where the implementations of biological functions are constrained by physical principles. Methods from statistical physics help to describe the functioning of complex biomolecular systems on a coarse-grained, but quantitative level.
Zellbiophysik (Prof. Andreas Bausch)
Understanding complex biomaterials on a fundamental physical basis is an integral challenge of future biophysical research. This challenge can be addressed by the concerted application of new experimental tools of soft condensed matter physics to living cells and bio-mimetic model systems. In our group we concentrate on the one hand on developing new physical tools to address the underlying complexity and mechanisms and on the other hand on developing new biomaterials for applications ranging from biomedicine to functional food.